Segment-swapped proteins offer a glimpse into the evolution of protein structures
|
A summary of our paper |
|
New paper in 2017: |
What is segment swapping?
There is a large diversity of basic protein architectures, or protein folds, in nature. The fundamental secondary structural elements (alpha helices and beta sheets) are combined into various compact structures such as multi-layer "sandwiches", "propellers", "barrels", "rolls", etc. How these diverse structures have evolved is an intriguing question.
While reviewing the data of a failed protein structure prediction experiment, we made an interesting observation. A protein having two consecutive domains was found to have a homolog that also had two domains but they were not consecutive; instead, the middle part of the chain formed one domain, and the two termini formed the other domain. We decided to find out whether there are other protein pairs with the same property. Sure enough, we found a number of such protein pairs. Realizing that we have discovered a novel group of proteins, we termed them segment-swapped proteins.
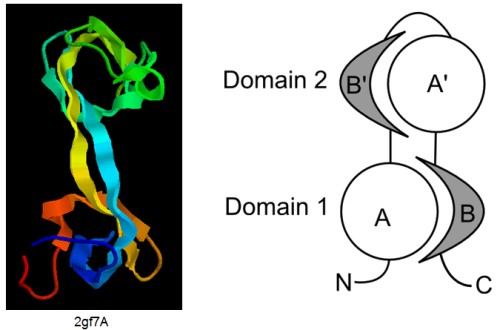
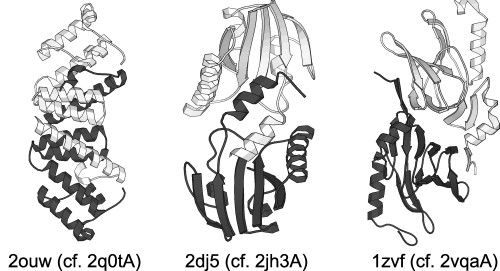
Segment-swapped proteins have two domains that are similar in structure, with one domain formed by the N- and C-termini (we call this Domain 1) and the other formed by the middle part of the chain (called Domain 2). In this architecture, Domain 2 consists of two segments that are structurally similar to the N- and C-terminal segments of Domain 1, but the order of the segments is reversed relative to Domain 1. If we denote the N- and C-terminal segments of Domain 1 by A and B then the entire chain can be represented as A-B'-A'-B, where A' and B' indicate segments structurally similar to A and B, respectively. A protein with two similar consecutive domains would be described as A-B-A'-B'.
Segment swapping is related to "3D domain swapping", a well-studied phenomenon. In 3D domain swapping, portions of proteins are swapped between two separate chains with identical sequences. In segment swapping, the swap occurs within a single chain. Because the sequences of the two halves of the chain are not identical but can diverge as the sequence evolves, the segment-swapped topology can become fixed during evolution.
The concept of segment swapping can be extended to multi-domain proteins. A three-domain segment-swapped protein would look like this:
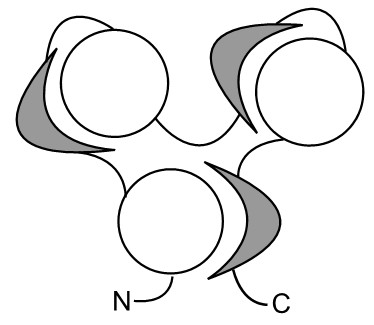
How many segment-swapped proteins are there in nature?
We developed a procedure to automatically recognize segment-swapped proteins by their structure, and scanned the Protein Data Bank to find all of them.
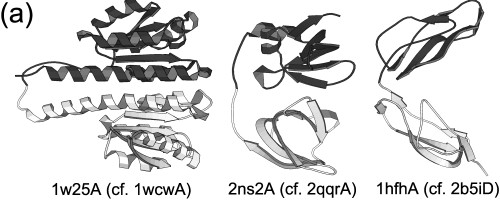
We identified 32 well-defined segment-swapped proteins belonging to 18 structural families. Segment-swapped proteins occur in all protein classes, and they are functionally diverse. This is a gallery of selected segment-swapped proteins:
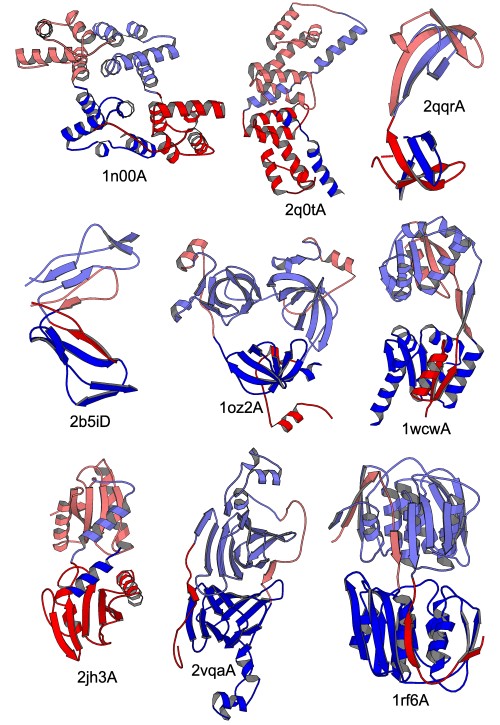
We even found a three-domain segment-swapped protein (1oz2A in the middle of the above gallery).
How have segment-swapped proteins evolved?
Segment-swapped proteins are extremely valuable for the study of the evolution of protein structures. We found that the two halves of their sequences usually have a relatively high sequence identity, which indicates that they are evolutionarily related, and are probably a result of gene duplication. We hypothesized two principal mechanisms by which segment-swapped proteins may have arisen in evolution:
- Domain swapping and fusion (DSF): The gene of an ancient single-domain protein duplicated and the two copies fused. The single-domain protein was capable of forming 3D domain swapped dimers, and it retained this ability after the gene duplication and fusion event.
- Circular permutation (CP): The gene of an ancient single-domain protein was subject to multiple gene duplication and fusion events. When three consecutive copies were present, the termini of this three-domain protein were truncated, and the remaining segments associated to form a new domain. The resulting sequence can be thought of as a circular permutation of a similar protein containing two consecutive domains.
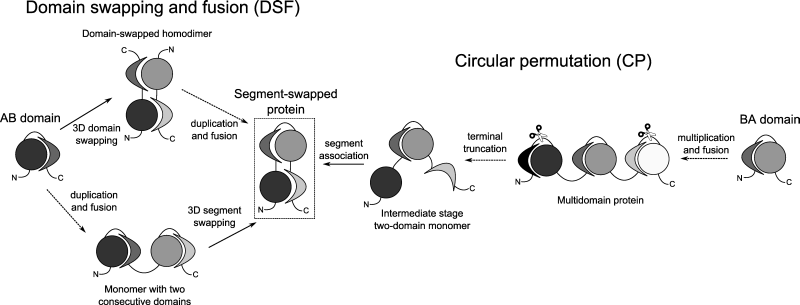
How can we decide which mechanism generated each segment-swapped proteins? This can be done by examining which of the two domains of the present-day protein is more ancient. If the protein is the result of the DSF mechanism, the ancient domain corresponds to Domain 1 of the present-day protein; but if it is the result of the CP mechanism, the ancient domain corresponds to Domain 2 of the present-day protein.
By performing structure and sequence comparisons between each domain of a segment-swapped protein and all the other structures in the Protein Data Bank, or complete genomes, we can obtain a rough estimate of which domain is more ancient.
Performing a number of such comparisons, we found that probably the DSF mechanism created most segment-swapped proteins. One exception is the 3-domain segment-swapped protein shown above, which was probably generated by the CP mechanism, similarly to beta propellers.
Consecutive versions of segment-swapped proteins
As expected, we can find some proteins that are consecutive versions of some segment-swapped proteins. Their sequences, however, differ from the segment-swapped versions.
Homodimeric analogs of segment-swapped proteins
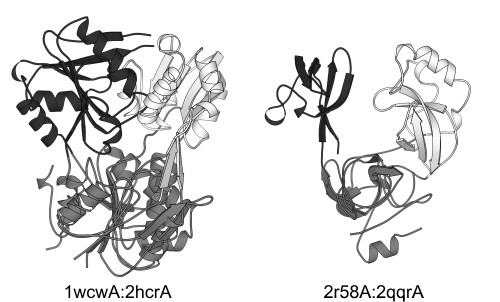
As mentioned, segment-swapping is in fact intra-chain 3D domain swapping. Thus, we can expect that some segment-swapped proteins have analogs that are 3D-domain-swapped homodimers. Indeed, we found a few examples for this:
Variants of segment-swapped proteins
Segment swapping can be thought of as starting with a protein having two consecutive domains, opening up both domains at some point, and rearranging the resulting segments by forming a new domain from the middle two segments and another from the two terminal segments. This implies that there may be some freedom regarding at which point we open up the domains to rearrange them (see Figure (a) below). Also, variants could also be generated by circular permutation (see Figure (b) below).
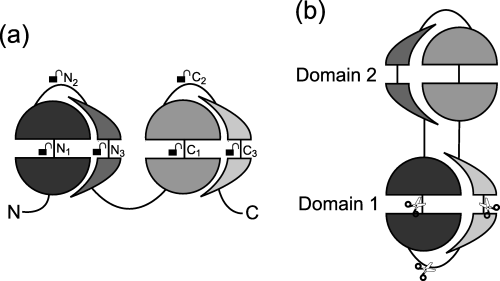
Indeed, we found some examples of protein pairs generated by opening up the domains at various sites as in Figure (a) above. The figures below show variants superimposed onto each other in their first domains which are shown in grey color, while the second domains are black and white, respectively.
We did not find any variants generated by circular permutation as in Figure (b) above.
The functional implications of segment swapping
Examining the functions of segment-swapped proteins, we found that they can be divided into two groups:
- The two domains work independently, each can bind some ligand (sometimes the substrate specificities may slightly differ).
- Ligands can bind into a cleft between the two domains, and a hinge-type relative motion of the domains is essential for function.
The majority of segment-swapped proteins belongs to the second group.
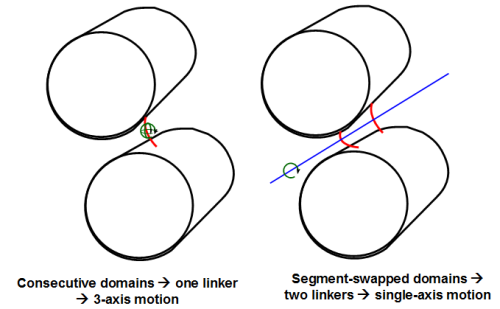
Is there an advantage to having segment-swapped rather than consecutive domains? Because in segment-swapped proteins, two peptide stretches connect the two domains rather than one, we hypothesize that having two connections makes it easier to develop a single-axis hinge-type motion, which has been shown to be functionally important in many proteins:
For more information on segment-swapped proteins, read our paper:
|
Szilágyi A, Zhang Y, Zavodszky P (2012): Intra-chain 3D segment swapping spawns the evolution of new multidomain protein architectures. J Mol Biol, 415(1):221-35. Pubmed PDF |